Angelika Böttger
Developmental signalling and apoptosis: Hydra vulgaris
 Hydra vulgaris is a member of the ancient phylum Cnidaria. We want to know how multicellular animals may have evolved from single celled protozoans and complex bilaterian animals from simply built non-bilaterians. Whole genome and EST sequencing of Hydra have shown that the major signalling pathways that are indispensable for development were already present in Cnidarians. The Notch signalling pathway is essential for cell specification and patterning in bilaterians. Similar processes are also influenced by Notch signalling in Hydra, e.g. stem cell differentiation and boundary formation. We have found interesting parallels in signalling events at the bud-parent boundary and boundaries that are formed during embryogenesis for organ formation in bilaterians. This might have implications for our understanding of the evolution of bilaterian body plans.
Hydra vulgaris is a member of the ancient phylum Cnidaria. We want to know how multicellular animals may have evolved from single celled protozoans and complex bilaterian animals from simply built non-bilaterians. Whole genome and EST sequencing of Hydra have shown that the major signalling pathways that are indispensable for development were already present in Cnidarians. The Notch signalling pathway is essential for cell specification and patterning in bilaterians. Similar processes are also influenced by Notch signalling in Hydra, e.g. stem cell differentiation and boundary formation. We have found interesting parallels in signalling events at the bud-parent boundary and boundaries that are formed during embryogenesis for organ formation in bilaterians. This might have implications for our understanding of the evolution of bilaterian body plans.
Multicellularity needs sophisticated mechanisms to maintain the equilibrium between different cell types within the organism. Apoptosis that is regulated by caspases and Bcl-2 family proteins is an important developmental mechanism to achieve such homeostasis in animals. We study the function of the surprisingly complex families of apoptotic proteins in Hydra in order to understand the evolutionary origins of apoptosis and how it is controlled by death and survival signals.
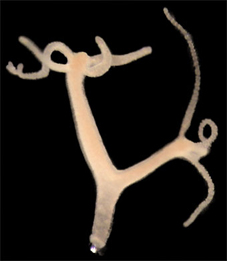
Picture: DAPT treated animal
Oxygen and gene expression: Jmjd6
 Research on a new Fe(II) and 2-oxo-glutarate dependent dioxygenase revealed that it has an impact on mRNA splicing. The protein Jmjd6 binds to several splice factors (e.g.U2AF-65). It hydroxylates its substrates on lysine residues. This results in changes in alternative splicing of some mRNAs. We are investigating the molecular mechanism that leads to these changes and the oxygen dependency of the splice process. We are trying to identify components of the spliceosome, that are modified by Jmjd6 and transcripts that are subject to splice regulation by Jmjd6. The importance of splice regulation by Jmjd6 is seen in Jmjd6 knockout mice and knockdown zebrafish, which have severe developmental defects and are not viable. We aim to understand the precise molecular function of Jmjd6 in development.
Research on a new Fe(II) and 2-oxo-glutarate dependent dioxygenase revealed that it has an impact on mRNA splicing. The protein Jmjd6 binds to several splice factors (e.g.U2AF-65). It hydroxylates its substrates on lysine residues. This results in changes in alternative splicing of some mRNAs. We are investigating the molecular mechanism that leads to these changes and the oxygen dependency of the splice process. We are trying to identify components of the spliceosome, that are modified by Jmjd6 and transcripts that are subject to splice regulation by Jmjd6. The importance of splice regulation by Jmjd6 is seen in Jmjd6 knockout mice and knockdown zebrafish, which have severe developmental defects and are not viable. We aim to understand the precise molecular function of Jmjd6 in development.
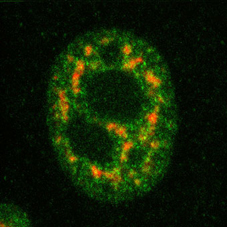
Picture: Endogenous Jmjd6 (green) and splice factor SC35 (red) in HeLa cells.

